Cell-based cluster model#
To illustrate an example from a cell model, we will use Model B from Reynolds et al. 2020, “Astrophysical Limits on Very Light Axion-like Particles from Chandra Grating Spectroscopy of NGC 1275”. It is an approximate model for a turbulent field in the Perseus cluster.
The total magnetic field strength at a distance \(z\) along the line of sight is set according to
where the electron density \(n_e\) is set by the analytic profile given by Churazov et al:
The cell sizes \(\Delta z\) are set according to a probability distribution function \(p(\Delta z)\), given by a power-law so that
spanning \(3.5-10\), kpc. In their model B, Reynolds et al. 2020 scale these minimum and maximum cell sizes linearly with radius as \(z/50{\rm kpc}\), because coherence lengths are expected to grow with distance from the cluster centre.
[1]:
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import alpro
alpro.util.set_default_plot_params()
First we initialise the survival class and set up the ALP parameters
[2]:
s = alpro.Survival("1275b")
s.init_model()
s.set_params(1e-12 * 1e-9, 1e-13)
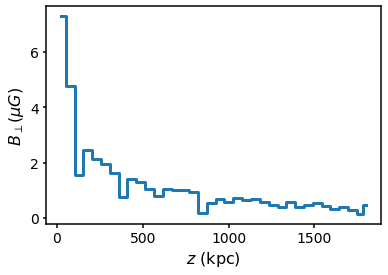
We can now initialise a random cell model with a given random number seed and plot \(B_\perp\) as a function of line of sight distance \(z\)
[3]:
Lmax = 1800.0
i = 0
s.domain.create_box_array(Lmax, i, s.coherence_func, r0=0)
plt.step(s.domain.rcen, s.domain.B * 1e6, where="mid")
plt.ylabel("$B_\perp (\mu G)$")
plt.xlabel("$z$ (kpc)")
[3]:
Text(0.5, 0, '$z$ (kpc)')
Now we can use ALPRO to compute the survival probability
[4]:
energies = np.logspace(3,4,1000)
P, Pradial = s.propagate(s.domain, energies, pol="both")
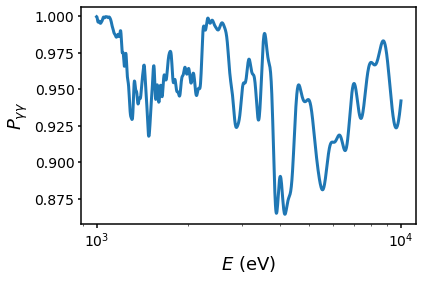
Let’s plot the survival probability first using the inbuilt method
[5]:
fig = s.default_plot(plot_kwargs = {"lw": 3}, mode="survival")
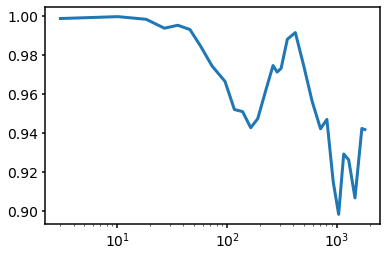
We can also plot the radial profile at the highest energy
[6]:
plt.plot(s.domain.rcen, 1.0 - Pradial[:,-1])
plt.semilogx()
[6]:
[]
We can also plot the radial profile in every energy bin using a colormesh plot
[7]:
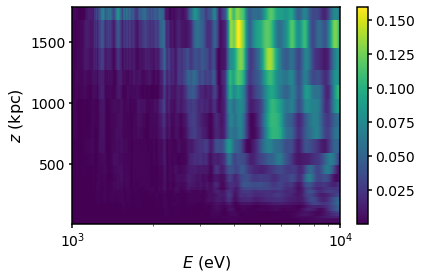
plt.pcolormesh(energies,s.domain.rcen, Pradial)
plt.colorbar()
plt.semilogx()
plt.xlabel("$E$ (eV)")
plt.ylabel("$z$ (kpc)")
[7]:
Text(0, 0.5, '$z$ (kpc)')
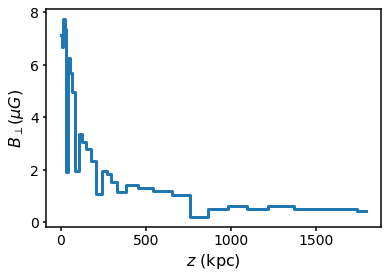
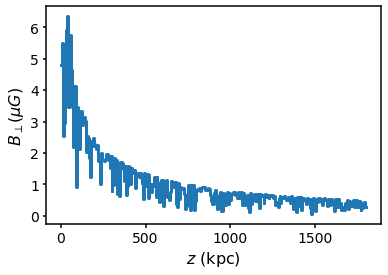
## Setting the coherence length / cell size distribution Model B uses a scaling of the coherence length with radius. This can be modified or turned off via the command set_coherence_r0. Setting it to None turns it off so the minimum and maximum scale lengths are constant
[8]:
s.set_coherence_r0(None)
s.domain.create_box_array(Lmax, i, s.coherence_func, r0=0)
plt.step(s.domain.rcen, s.domain.B * 1e6, where="mid")
plt.ylabel("$B_\perp (\mu G)$")
plt.xlabel("$z$ (kpc)")
[8]:
Text(0.5, 0, '$z$ (kpc)')
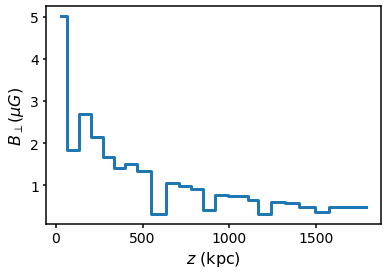
The power-law function can also be customised using the set_coherence_pl function. For example, if we want a PDF of the form \(p(\Delta z) \propto \Delta z^{-3}\) chosen between 50 and 100 kpc, we can do the following:
[9]:
s.set_coherence_pl(n=3, xmin=50.0, xmax=100.0, r0=None)
s.domain.create_box_array(Lmax, i, s.coherence_func, r0=0)
plt.step(s.domain.rcen, s.domain.B * 1e6, where="mid")
plt.ylabel("$B_\perp (\mu G)$")
plt.xlabel("$z$ (kpc)")
[9]:
Text(0.5, 0, '$z$ (kpc)')
For more control over the distribution of cell sizes, any function that chooses random numbers can be passed to the coherence_func object - a convenient way to do this is using a scipy stats function like rayleigh using the inbuilt rvs method.
[10]:
from scipy.stats import rayleigh
s.coherence_func = rayleigh(50).rvs
s.domain.create_box_array(Lmax, i, s.coherence_func, r0=0)
plt.step(s.domain.rcen, s.domain.B * 1e6, where="mid")
plt.ylabel("$B_\perp (\mu G)$")
plt.xlabel("$z$ (kpc)")
[10]:
Text(0.5, 0, '$z$ (kpc)')